ProteinMCP
An Agentic AI Framework for Autonomous Protein Engineering
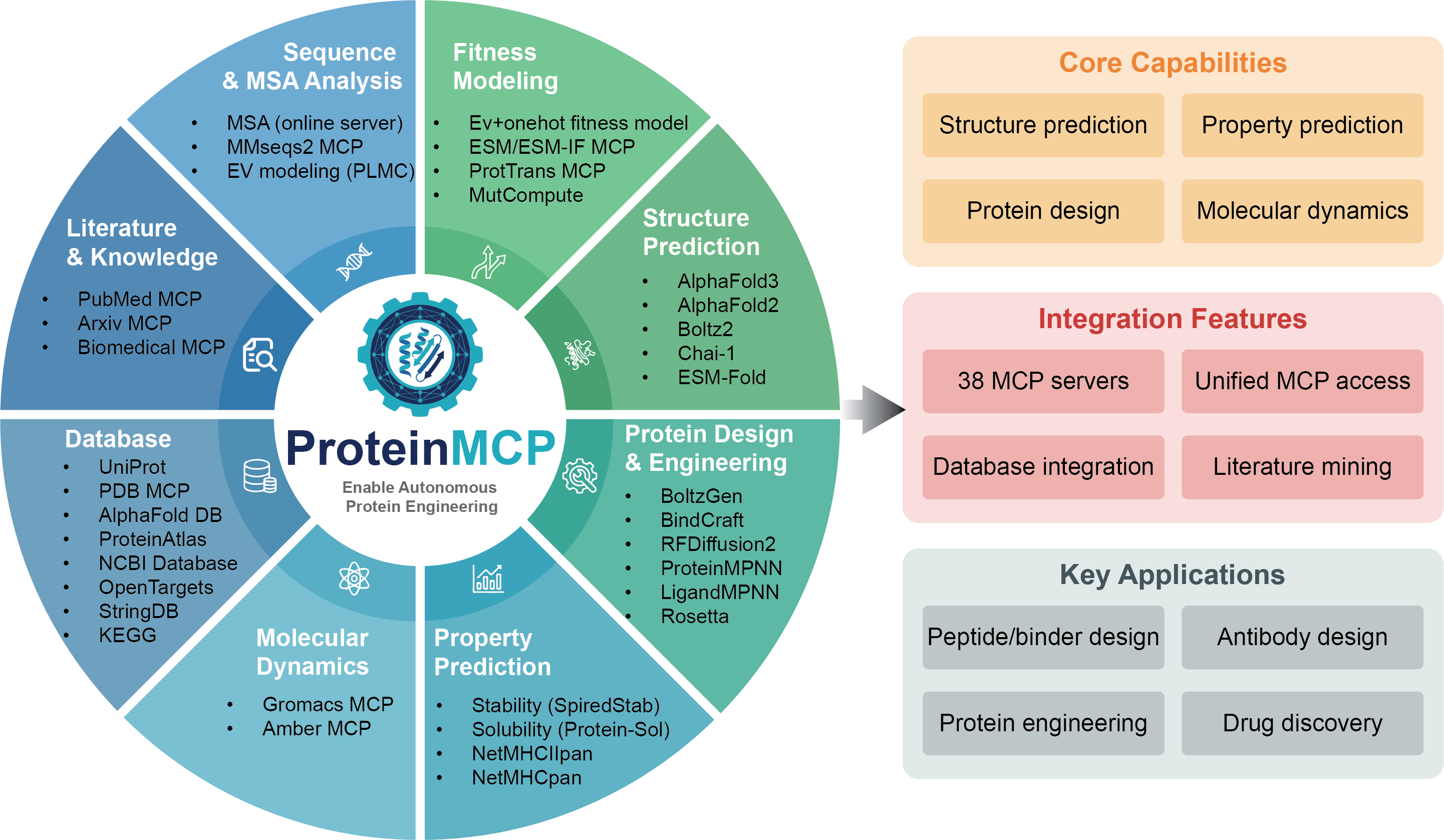
What is ProteinMCP?
ProteinMCP provides a registry of 38 MCP (Model Context Protocol) servers for protein engineering tools, a workflow skill system that orchestrates multi-MCP pipelines, and CLI tools (pmcp, pskill) to manage everything.
It enables Claude Code to autonomously execute complex protein engineering workflows — from fitness prediction to de novo binder design — by connecting AI agents to specialized computational biology tools.
Key Features
- 38 MCP Servers — Structure prediction, protein design, binder design, molecular dynamics, fitness modeling, sequence analysis, and immunology tools
- Workflow Skills — Guided multi-step pipelines that orchestrate multiple MCPs via Claude Code
- Two Runtime Types — Python (local venv) and Docker (GPU containers) for flexible deployment
- CLI Management —
pmcpandpskillcommands for installing, registering, and managing tools - Auto MCP Creation — Generate new MCP servers from any GitHub repository with
pmcp create
Demo Workflows
Protein Fitness Prediction
Predict the fitness landscape of protein variants using multiple ML approaches. The workflow generates MSA, builds evolutionary coupling models, trains ESM and ProtTrans embeddings, and compares all models with publication-ready visualizations.
Required MCPs: msa_mcp, plmc_mcp, ev_onehot_mcp, esm_mcp, prottrans_mcp
De Novo Binder Design
Design protein binders against a target using BindCraft’s integrated pipeline of RFdiffusion backbone generation, ProteinMPNN sequence design, and AlphaFold2 validation.
Required MCPs: bindcraft_mcp
Nanobody CDR Design
Design nanobody CDR loop regions targeting a protein using BoltzGen with specialized nanobody protocols and cysteine filtering.
Required MCPs: boltzgen_mcp
Quick Example
# Install ProteinMCP
mamba env create -f environment.yml
mamba activate protein-mcp
pip install -e .
# Install a workflow skill (auto-installs all required MCPs)
pskill install fitness_modeling
# Launch Claude Code and run the skill
claude
> /fitness-model
Claude will prompt you for inputs (protein name, data location, etc.) and execute the full pipeline autonomously.